This function gets the standards data and plots the standard curves for any antigens (i.e., non-PvSeroTaT specific).
Value
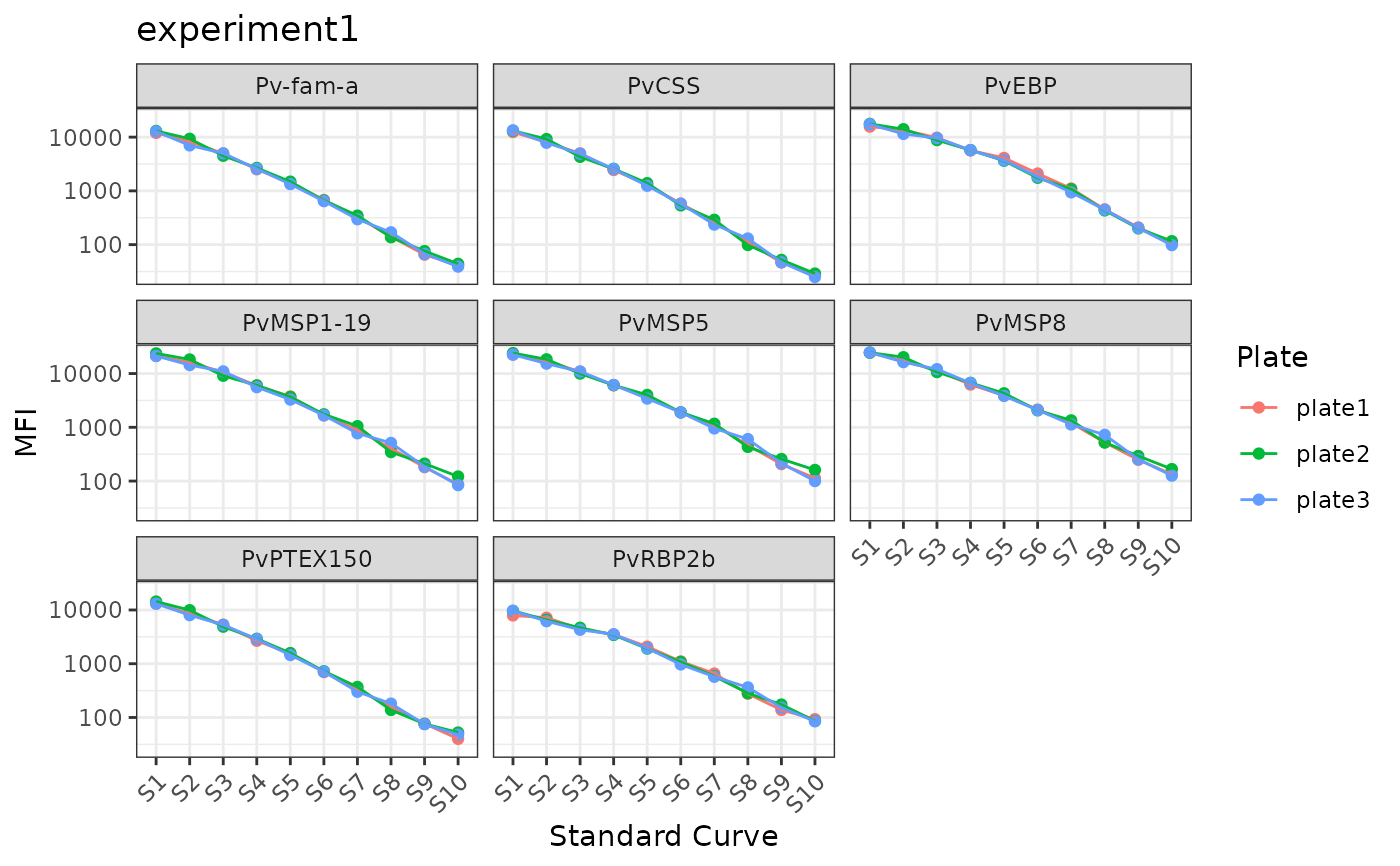
- Dot and line plot of standard curves (S1-S10) - WEHI-acceptable standard curve data on background of plot with user data.
Examples
# \donttest{
# Example demonstrating how to process bead count data.
# These files are included in the SeroTrackR package under inst/extdata.
your_raw_data <- c(
system.file("extdata", "example_MAGPIX_plate1.csv", package = "SeroTrackR"),
system.file("extdata", "example_MAGPIX_plate2.csv", package = "SeroTrackR"),
system.file("extdata", "example_MAGPIX_plate3.csv", package = "SeroTrackR")
)
# Read in raw MAGPIX data
sero_data <- readSeroData(
raw_data = your_raw_data,
platform = "magpix"
)
#> PASS: File example_magpix_plate1.csv successfully validated.
#> PASS: File example_magpix_plate2.csv successfully validated.
#> PASS: File example_magpix_plate3.csv successfully validated.
# Plot Standards
plotStds_all(
sero_data = sero_data,
experiment_name = "experiment1"
)
 # }
# }
