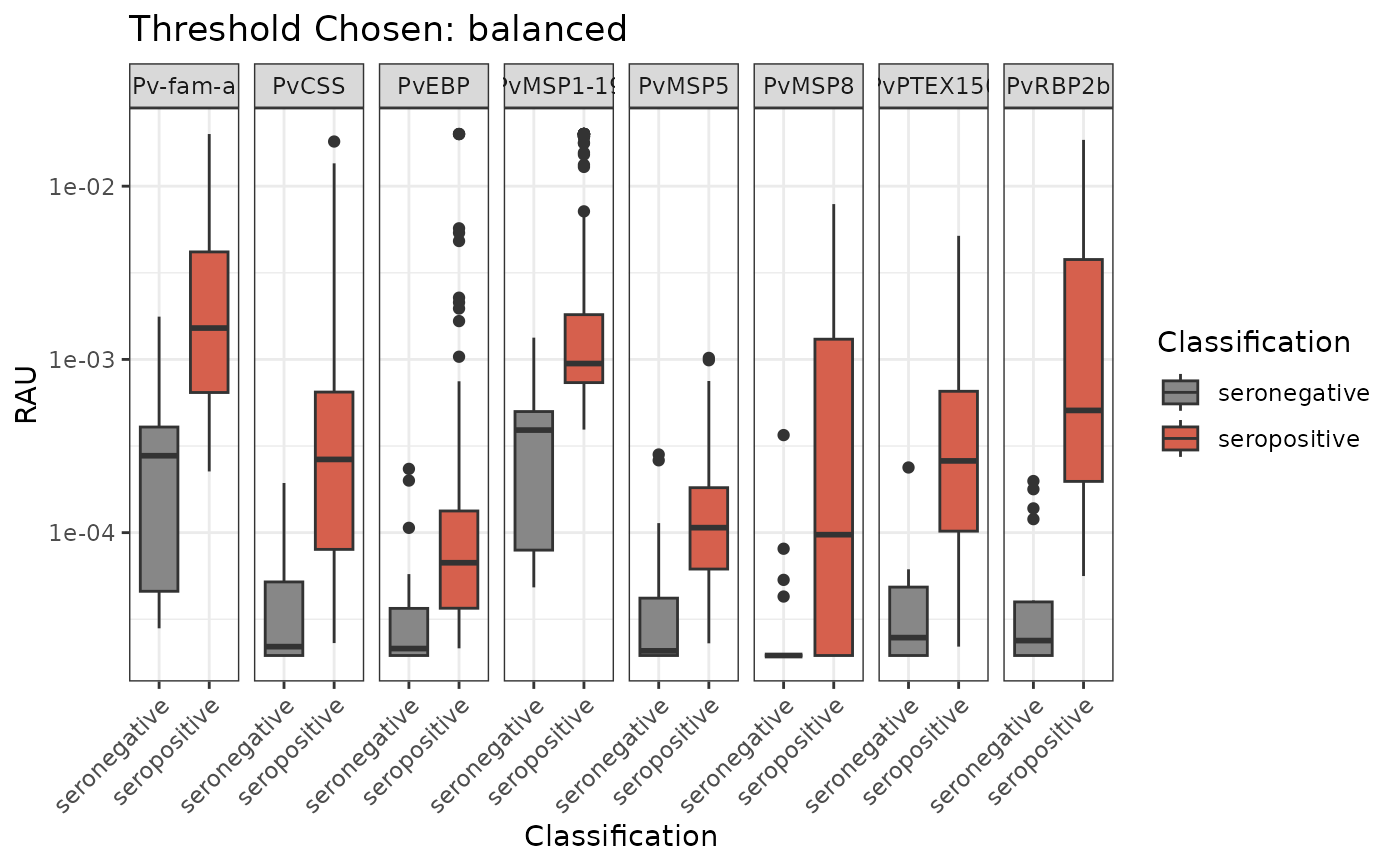
One example of data visualisation to detect the median and interquartile range of the RAU values per antigen for seropositive and seronegative individuals. Please note that the `classifyResults()` function must be run first.
Examples
# \donttest{
# Step 0: Load example raw data
your_raw_data <- c(
system.file("extdata", "example_MAGPIX_plate1.csv", package = "SeroTrackR"),
system.file("extdata", "example_MAGPIX_plate2.csv", package = "SeroTrackR")
)
your_plate_layout <- system.file(
"extdata",
"example_platelayout_1.xlsx",
package = "SeroTrackR"
)
# Step 1: Read serology data and plate layout
sero_data <- readSeroData(your_raw_data,"magpix")
#> PASS: File example_magpix_plate1.csv successfully validated.
#> PASS: File example_magpix_plate2.csv successfully validated.
plate_list <- readPlateLayout(your_plate_layout, sero_data)
#> Plate layouts correctly identified!
# Step 2: Process counts and perform quality control
qc_results <- runQC(sero_data, plate_list)
# Step 3: Convert MFI to RAU using ETH beads
mfi_to_rau <- MFItoRAU_Adj(
sero_data = sero_data,
plate_list = plate_list,
qc_results = qc_results
)
#> Joining with `by = join_by(antigen)`
#> Joining with `by = join_by(antigen)`
#> Joining with `by = join_by(antigen)`
#> Joining with `by = join_by(antigen)`
#> Joining with `by = join_by(antigen)`
#> Joining with `by = join_by(antigen)`
# Step 4: Define sens/spec thresholds
sens_spec_all <- c(
"balanced", "85% sensitivity", "90% sensitivity", "95% sensitivity",
"85% specificity", "90% specificity", "95% specificity"
)
# Step 5: Classify results across all thresholds
all_classifications <- purrr::map_dfr(sens_spec_all, ~{
classifyResults(
mfi_to_rau_output = mfi_to_rau,
algorithm_type = "antibody_model",
sens_spec = .x,
qc_results = qc_results
) |>
as.data.frame() |>
dplyr::mutate(sens_spec = .x)
})
# Plot classification for a single threshold
plotBoxPlotClassification(all_classifications, "balanced")
 # }
# }
